Analysis of Impact of Data size for Classification of Alzheimer’s disease using Convolution neural network
Keywords – Deep learning, Alzheimer’s Disease, Convolution Neural Network, PET
I. INTRODUCTION
Alzheimer’s Disease (AD) is a type of neurodegenerative disorder and is a common type of Dementia that accounts for 70 to 80 percent of Dementia cases [1]. It is characterized by the decline in thinking ability and changes in behavior starting in middle to old age. The symptoms develop gradually but degrade over time to a point where it can get serious enough to interfere with normal life. In the early stages, there is a mild memory loss while in the later stages, the patient’s ability to respond and hold a conversation degrades dramatically[2]. As the disease progresses, symptoms include difficulties with speech and writing, problem-solving, decreased judgment among others. During the advanced stages of AD, the neural damage hampers normal bodily functioning such as basic motor functions for instance walking, talking etc., and during the final stages, the patient becomes completely dependent upon their caregivers.
AD affects nearly 45 million worldwide and these numbers are expected to increase many-fold in the coming decades and is projected to grow to nearly 90 million by the year 2050. According to 2018 Alzheimer’s Disease Facts and Figures[1], an estimated 6 million Americans are suffering from Alzheimer’s including an estimated 5.5 million above the age of 65 and another 200,000 under 65 who have a young-onset of Alzheimer’s disease. AD is one of the main causes of Death in the US and is the only disease in the top 10 leading causes of death that cannot be prevented. Whilst the number of death cases due to other ailments such as Heart disease, Cancer, Diabetes etc., are on a decline, the number of deaths due to AD have increased by about 120% in the past 15 years.
The truth is that only a fraction of the people with AD is recognized in the primary care[3]. Brain imaging techniques, namely Single Photon Emission Computed Tomography (SPECT), Magnetic resonance imaging (MRI) and Positron Emission Tomography (PET) have shown to be a very effective method for diagnosis of AD using methods that are non-invasive. Early detection of Alzheimer’s disease has a lot of benefits that can help the patient and their loved ones in making timely decisions. Early diagnosis allows people with AD and their clos ones to receive proper advice, and support; giving them access to the best drug and non-drug therapies that can improve their perception and enhance their quality of life. Undetected Alzheimer’s disease places them at risk of hallucinations, accidents, medication errors, and financial difficulties etc. Hence, this project aims at improving the accuracy of early detection of Alzheimer’s disease in order to help people take right decision at the right time.
With this in mind, in 2004, the Alzheimer’s Disease Neuroimaging Initiative (ADNI) was launched, funded by National Institute on Aging (NIA), the Foundation for the National Institutes of Health, and other companies as a public-private partnership with the goal of analysing whether a combination of PET, MRI, SPECT or other biomarkers and clinical assessments will be useful for detecting AD at an early stage. Several researchers, taking advantage of the large dataset available through ADNI have developed several advanced ML and pattern recognition models to map the degenerative patterns and identify those who are at a peril of AD.
II. LITERATURE SURVEY
Various classification methods have been proposed to automatically classify between affected patients and healthy individuals. Examples of few techniques used for classification include support vector machine (SVM)[4], boosting[5], artificial neural networks[6], k-nearest neighbor[7] and linear discriminant analysis[8]. There are two broad approaches to classifying the data;
i. Single Modal Approach and,
ii. Multimodal Approach.
Also, there are two main learning algorithms used for the process of classification viz., machine learning techniques such as Support vector machine, k-nearest neighbor, Linear regression etc., and the other using deep learning algorithms such as Convolution neural network. Whilst Machine Learning is a statistical model of learning in which a dataset is described using a features-list, Deep Learning on the other hand uses statistical learning to extract features or attributes from raw data.
i. SINGLE MODALITY APPROACHES
In the following, we will briefly discuss a few approaches used by researchers. Stefan Klöppel, Cynthia M. Stonnington et al. (2008), the dataset collected was divided into 4 groups 1,2,3 and 4 consisting of 20 AD and 20 Control normal, 14 AD and 14 Control normal, 33 with mild AD and 57 healthy patients and, the fourth group consisted of 18 AD patients and 19 individuals with confirmed FTLD respectively. For the first and third groups, MRI scans over a period of 10 years were considered. Images were inspected for artifacts and abnormalities through visual means. They were then sectioned off into grey matter (GM), white matter and cerebrospinal fluid and they were trained with Support Vector Machine Algorithm[3]. For effective classification, the input datasets used were pathologically proven and were obtained from different medical centers[2]. The researchers were able to accurately classify 96% of the patients.
In 2010, R. Chaves, J. Ramírez et al. collected the SPECT data from 97 participants out of which 54 suffered from AD while the rest 43 were normal controls. The authors, using Association Rule Mining, determined the association between the features in the datasets using activated brain regions of interest. These regions of interest were obtained using voxels of each image and considering them as features. The results showed a classification accuracy of 95.87%. Saman Sarraf et al. (2016) used Convolution neural network on the fMRI data of 28 AD patients and 15 normal controls. The dataset was separated into three sections: 60% for training, and 20% each for both validation and testing. For training, 30 epochs were selected along with a batch size of 64, yielding 126,990 iterations[13]. The authors were able to successfully classify AD from normal with an accuracy of 96.86%. To our best information, this is the highest accuracy attained for a single modality classification to date.
ii. MULTIMODAL APPROACH
While the use of individual imaging technique has yielded favourable results, these models are designed to describe the differences but this cannot be used to extend to the individual. With this in mind, R Chaves et al. (2013) used the SPECT data of 41 Control Normal and 56 AD patients divided into 3 groups of 30, 22 and 4 patients along with PET data of 75 each AD and Control normal. Classification system consisted of four stages: (i) procedural masking, (ii) mean intensity discretization, (iii) AR mining, (iv) cross-validation with leave-one-out (loo)[11]. The authors were able to successfully classify SPECT data with an accuracy of 96.91% while the accuracy achieved for just PET was 92%.
Zhang et al.. (2011) proposed a method to classify AD patients from healthy ones using three different biomarkers i.e., CSF, PET and MRI. The authors used the baseline data set with a total of 202 subjects, of which 51 had Alzheimer’s, 99 had MCI and 52 had no conditions and were considered as healthy. Different tests were conducted for MRI, PET and CSF and an accuracy of 93.2% was achieved. Authors claimed that multimodal classification method using the three imaging techniques i.e., PET, CSF and MRI yields better results and is more robust over those using only one imaging technique, for any of number of regions in the brain selected[2].
iii. LIMITATIONS
These studies we outlined are but a few examples of the strides being made and obviously, there are a lot many other impressive studies with good results. But that cannot let us turn a blind eye to the potential issues in the process. Being able to identify issues whether it be with the input data, algorithm, validation etc., is an important part of the prognosis. With that in mind, some of the main issues we found include issues such as sample size, class imbalance, and usage of the unproven dataset. While it is easy to achieve higher accuracies, these methods cannot be used to characterize a larger population of data. Also, small samples are prone to overtraining. Imbalanced classes and usage of unproven data along with the usage of small data sizes put some doubt on the robustness of the training algorithms.
III. METHODOLOGY AND ALGORITHM
While Machine learning algorithms are useful for applications where the data sizes are small to medium, Deep learning algorithms benefit from larger dataset. Going with the hypothesis that larger datasets improve the robustness of the classification process, we will be going with a deep learning algorithm known as Convolution Neural Network (CNN). The process of initializing a Convolutional Neural Network involves four major steps:
Step I : Convolution layer
Step II : Pooling layers
Step III : Flattening the pooled output
Step IV : Full connection of the flattened neurons

Figure 1. Workflow of CNN
We acquired the PET data of 95 patients with confirmed AD and 104 healthy individuals from ADNI. The network was initialized as a sequential network using the Keras image processing library and the Keras layer for convolution was used for training and testing. Also, we used the Maxpooling function for pooling. In Maxpooling, the pixel with the maximum value from the selected region of interest is needed. The next layers flatten and dense convert the 2D matrix into a single-dimensional continuous linear vector and perform the full connection of the network respectively. The convolution function generates the convolution layers based on the filters count, its shape, shape of the input image and the type of image. Here, we are using three convolution layers with the first two containing 32 filters and the last one containing 64 filters, all 3*3 in size. The images were monochromatic and of size 128*128 pixels. The activation function ‘relu’ was used.
Maxpooling with a pooling matrix of 2*2 was used to get the precise region of the features and have a minimum pixel loss. The primary aim of pooling is to condense the size of the image to a smaller size as much as possible. The key thing to understand here is to reduce the total number of nodes for the upcoming layers. The images were pre-processed to avoid over-fitting. Overfitting takes place when the accuracy for training is great but the test accuracy is extremely low due to overfitting of nodes between the different consecutive layers. The images were given to the model for training and testing across 10 epochs and the final result was stored in an HDF5 file for prediction testing.
We divided the data into five groups; 1, 2, 3, 4 and 5; for training, and testing starting from a small group consisting of 10 AD patients and 10 healthy individuals to the whole dataset with 95 AD patients and 104 normal controls. In order to maintain robustness each data set was trained and tested across five runs and the mean values of these were considered. All the groups were divided in a ratio of approximately nine-part for training to one part for validation.
For the first group, we considered a dataset of 10 patients suffering from AD and 10 patients who were tested to be healthy. With this, we divided the group to 9 people for training and one person from each for testing. Next, we considered a group of 25 AD patients and 25 control normal divided into 22 each for training and 3 subjects for testing. The third group consisted of 50 each from AD and healthy individuals which we dived into 45 subjects for training and 5 for validation. The fourth group consisted of 75 individuals each who were separated into 67 each for training the algorithm and 8 each for testing and the final group consisted of the whole data set where 85 patients each were used for training and the rest for from each groups were used for testing.
IV. RESULTS AND DISCUSSIONS
A detailed analysis of the result is given herewith. The main goal of using small test cases was to showcase the issue of overfitting when using a small dataset. When a small sample size is used in the training process, the neural network tends to “memorize” rather than learn; leading to a very high training accuracy but a low validation accuracy. This result can be seen in the first two groups very prominently where the mean training accuracies are 99.6% and 95.2% respectively but the validation accuracy were merely 46.34% and 50.61% respectively.
TABLE I. Training accuracies achieved in 5 runs
| Group 1 (10 AD, 10 CN) | |||||
| T1 | T2 | T3 | T4 | T5 | Mean |
| 99.60 | 99.62 | 99.55 | 99.63 | 99.64 | 99.61 |
| Group 2 (25 AD, 25 CN) | |||||
| T1 | T2 | T3 | T4 | T5 | Mean |
| 95.22 | 95.19 | 95.13 | 95.25 | 95.26 | 95.21 |
| Group 3 (50 AD, 50 CN) | |||||
| T1 | T2 | T3 | T4 | T5 | Mean |
| 92.11 | 91.97 | 91.96 | 92.00 | 91.99 | 92.01 |
| Group 4 (75 AD, 75 CN) | |||||
| T1 | T2 | T3 | T4 | T5 | Mean |
| 87.27 | 87.34 | 87.33 | 87.30 | 87.29 | 87.31 |
| Group 5 (95 AD, 104 CN) | |||||
| T1 | T2 | T3 | T4 | T5 | Mean |
| 82.85 | 82.80 | 82.86 | 82.82 | 82.83 | 82.83 |

Figure. 2. Training accuracies (Mean) of 5 runs
TABLE II. Validation accuracies achieved in 5 runs
| Group 1 (10 AD, 10 CN) | |||||
| T1 | T2 | T3 | T4 | T5 | Mean |
| 46.33 | 46.38 | 46.31 | 46.32 | 46.35 | 46.34 |
| Group 2 (25 AD, 25 CN) | |||||
| T1 | T2 | T3 | T4 | T5 | Mean |
| 50.60 | 50.64 | 50.61 | 50.61 | 50.58 | 50.61 |
| Group 3 (50 AD, 50 CN) | |||||
| T1 | T2 | T3 | T4 | T5 | Mean |
| 55.69 | 55.61 | 55.63 | 55.64 | 55.63 | 55.64 |
| Group 4 (75 AD, 75 CN) | |||||
| T1 | T2 | T3 | T4 | T5 | Mean |
| 59.29 | 59.35 | 59.31 | 59.28 | 59.30 | 59.31 |
| Group 5 (95 AD, 104 CN) | |||||
| Run 1 | Run 2 | Run 3 | Run 4 | Run 5 | Mean |
| 64.30 | 64.36 | 64.31 | 64.31 | 64.29 | 64.31 |
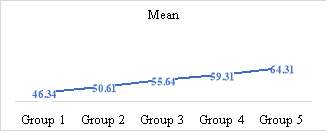
Figure 3. Validation accuracies (Mean ) of 5 runs
With the group 3 and 4, the mean accuracies did drop with accuracies of 92% and 87.31% respectively but the validation was on an upward trend with 55.64% and 59.31% respectively and the last group with the whole dataset achieved an accuracy of 82.83% with a validation accuracy of 64.31%. The results showcase the importance of having a large, class balanced data set for improved prediction accuracy. As the data size increased the overall training accuracy decreased. This downward trend in the accuracy can be attributed to the presence of abnormalities in the training data. With further optimizations in the learning algortihm we will be able to attain better training accuracies than those achieved here, but that process is out of the scope of this research. The main take away point from this is that the validation accuracy increases with the increase in sample size and that machine learning and deep learning techinques do indeed greatly benefit from large datasets.
For the Alzheimer’s Disease Neuroimaging Initiative*
*Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at:
http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf
REFERENCES
[1] Alzheimer’s Association, “Alzheimer’s Disease Facts and Figures” Alzheimer’s and Dementia 2018, pp. 367-429
[2] Aunsia Khan, Muhammad Usman “Early Diagnosis of Alzheimer’s Disease using Machine Learning Techniques” 7th International Joint Conference on Knowledge Discovery, Knowledge Engineering and Knowledge Management (IC3K), 2015, pp. 380-388
[3] Klöppel, S., C. M. Stonnington, et al. Automatic classification of MR scans in Alzheimer’s disease. Brain 2008, pp. 681-689
[6] Vapnik VN The nature of statistical learning theory, 1999, Springer-Verlag
[7] Morra JH, Tu Z, Apostolova LG, et al., “Comparison of AdaBoost and support vector machines for detecting Alzheimer’s Disease through automated hippocampal segmentation.” IEEE T Med Imaging, 2010, pp. 30-43
[8] Jiang J, Trundle P et al “Medical image analysis with artificial neural networks.” Computerized Medical Imaging and Graphics 2010, pp. 617-631
[9] Fitzpatrick JM, Sonka M Handbook of Medical Imaging, Volume 2. SPIE-The International Society for Optical Engineering, 2000
[9] Bankman Handbook of Medical Image Processing and Analysis, Academic Press, 2008
[10] Chaves, R., J. Ramírez, et al., “Effective Diagnosis of Alzheimer’s Disease by Means of Association Rules. Hybrid Artificial Intelligence Systems”, Springer, 2010, pp. 452-459
[11] Chaves, R., J. Ramírez, et al., “Integrating discretization and association rule-based classification for Alzheimer’s disease diagnosis”, 2013, pp. 1571-1578
[12] Chong-Yaw Wee, Daoqiang Zhang, Luping Zhou et al., “Machine Learning Techniques for AD/MCI Diagnosis and Prognosis”, 2013, pp. 147-179
[13] Saman Sarraf, Ghassem Tofighi, “Deep Learning-based Pipeline to Recognize Alzheimer’s Disease using fMRI Data”, 2016, pp. 816-820
[14] D, Zhang. “Multimodal classification of Alzheimer’s disease and mild cognitive impairment.” Neuroimage, 2011, pp. 856-867.
Cite This Work
To export a reference to this article please select a referencing style below:
Related Content
All TagsContent relating to: "analysis"
Analysis is the process of breaking a complex topic or substance into smaller parts in order to gain a better understanding of it. In the nursing process this often forms part of a framework for the essential components of nursing practice. These steps are assessment and data gathering, analysis, planning, intervention, and evaluation.
Related Articles


